Published: 21 March 2022, MDPI
A Deep Learning Model for Prostate Adenocarcinoma Classification in Needle Biopsy Whole-Slide Images Using Transfer Learning
The histopathological diagnosis of prostate adenocarcinoma in needle biopsy specimens is of pivotal importance for determining optimum prostate cancer treatment. Since diagnosing a large number of cases containing 12 core biopsy specimens by pathologists using a microscope is time-consuming manual system and limited in terms of human resources, it is necessary to develop new techniques that can rapidly and accurately screen large numbers of histopathological prostate needle biopsy specimens. Computational pathology applications that can assist pathologists in detecting and classifying prostate adenocarcinoma from whole-slide images (WSIs) would be of great benefit for routine pathological practice. In this paper, we trained deep learning models capable of classifying needle biopsy WSIs into adenocarcinoma and benign (non-neoplastic) lesions. We evaluated the models on needle biopsy, transurethral resection of the prostate (TUR-P), and The Cancer Genome Atlas (TCGA) public dataset test sets, achieving an ROC-AUC up to 0.978 in needle biopsy test sets and up to 0.9873 in TCGA test sets for adenocarcinoma. Read full text
Published: 24 February 2022, MDPI
A deep learning model for cervical cancer screening on liquid-based cytology specimens in whole slide images
Liquid-based cytology (LBC) for cervical cancer screening is now more common than the conventional smears, which when digitised from glass slides into whole-slide images (WSIs), opens up the possibility of artificial intelligence (AI)-based automated image analysis. Since conventional screening processes by cytoscreeners and cytopathologists using microscopes is limited in terms of human resources, it is important to develop new computational techniques that can automatically and rapidly diagnose a large amount of specimens without delay, which would be of great benefit for clinical laboratories and hospitals. The goal of this study was to investigate the use of a deep learning model for the classification of WSIs of LBC specimens into neoplastic and non-neoplastic. To do so, we used a dataset of 1605 cervical WSIs. We evaluated the model on three test sets with a combined total of 1468 WSIs, achieving ROC AUCs for WSI diagnosis in the range of 0.89–0.96, demonstrating the promising potential use of such models for aiding screening processes. Read full text
Published: 25 January 2022, Springer Nature
A deep learning model for breast ductal carcinoma in situ classification in whole slide images
The pathological differential diagnosis between breast ductal carcinoma in situ (DCIS) and invasive ductal carcinoma (IDC) is of pivotal importance for determining optimum cancer treatment(s) and clinical outcomes. Since conventional diagnosis by pathologists using microscopes is limited in terms of human resources, it is necessary to develop new techniques that can rapidly and accurately diagnose large numbers of histopathological specimens. Computational pathology tools which can assist pathologists in detecting and classifying DCIS and IDC from whole slide images (WSIs) would be of great benefit for routine pathological diagnosis. In this paper, we trained deep learning models capable of classifying biopsy and surgical histopathological WSIs into DCIS, IDC, and benign. We evaluated the models on two independent test sets (n= 1382, n= 548), achieving ROC areas under the curves (AUCs) up to 0.960 and 0.977 for DCIS and IDC, respectively. Read full text
Published: 9 November 2021, MDPI
Deep learning models for poorly differentiated colorectal adenocarcinoma classification in whole slide images using transfer learning
Colorectal poorly differentiated adenocarcinoma (ADC) is known to have a poor prognosis as compared with well to moderately differentiated ADC. The frequency of poorly differentiated ADC is relatively low (usually less than 5% among colorectal carcinomas). Histopathological diagnosis based on endoscopic biopsy specimens is currently the most cost effective method to perform as part of colonoscopic screening in average risk patients, and it is an area that could benefit from AI-based tools to aid pathologists in their clinical workflows. In this study, we trained deep learning models to classify poorly differentiated colorectal ADC from Whole Slide Images (WSIs) using a simple transfer learning method. We evaluated the models on a combination of test sets obtained from five distinct sources, achieving receiver operating characteristic curve (ROC) area under the curves (AUCs) up to 0.95 on 1799 test cases. Read full text
Published: 26 October 2021, MDPI
Breast invasive ductal carcinoma classification on whole slide images with weakly-supervised and transfer learning
Invasive ductal carcinoma (IDC) is the most common form of breast cancer. For the non-operative diagnosis of breast carcinoma, core needle biopsy has been widely used in recent years for the evaluation of histopathological features, as it can provide a definitive diagnosis between IDC and benign lesion (e.g., fibroadenoma), and it is cost effective. Due to its widespread use, it could potentially benefit from the use of AI-based tools to aid pathologists in their pathological diagnosis workflows. In this paper, we trained invasive ductal carcinoma (IDC) whole slide image (WSI) classification models using transfer learning and weakly-supervised learning. We evaluated the models on a core needle biopsy (n = 522) test set as well as three surgical test sets (n = 1129) obtaining ROC AUCs in the range of 0.95–0.98. The promising results demonstrate the potential of applying such models as diagnostic aid tools for pathologists in clinical practice. Read full text
Published: 14 October 2021, Scientific Reports
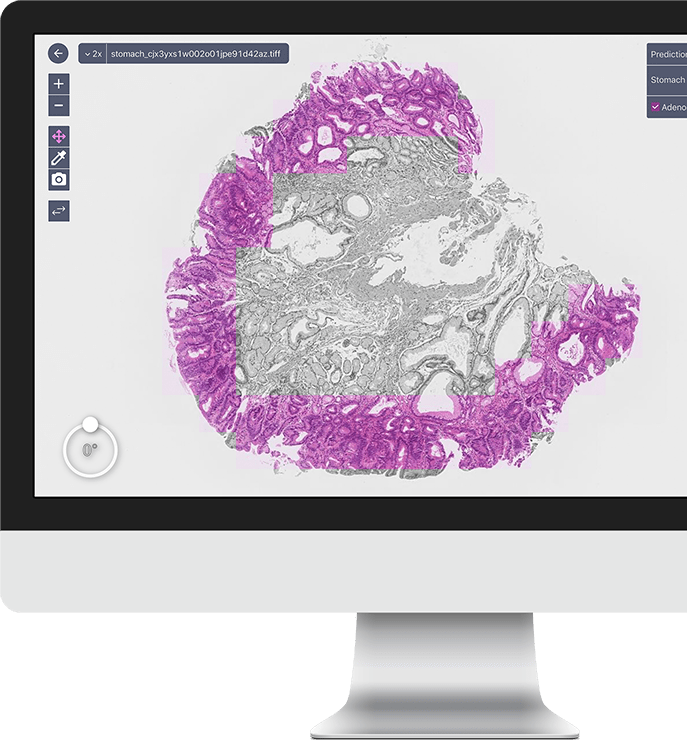
A deep learning model for gastric diffuse-type adenocarcinoma classification in whole slide images
Gastric diffuse-type adenocarcinoma represents a disproportionately high percentage of cases of gastric cancers occurring in the young, and its relative incidence seems to be on the rise. Usually it affects the body of the stomach, and it presents shorter duration and worse prognosis compared with the differentiated (intestinal) type adenocarcinoma. The main difficulty encountered in the differential diagnosis of gastric adenocarcinomas occurs with the diffuse-type. As the cancer cells of diffuse-type adenocarcinoma are often single and inconspicuous in a background desmoplaia and inflammation, it can often be mistaken for a wide variety of non-neoplastic lesions including gastritis or reactive endothelial cells seen in granulation tissue. In this study we trained deep learning models to classify gastric diffuse-type adenocarcinoma from WSIs. We evaluated the models on five test sets obtained from distinct sources, achieving receiver operator curve (ROC) area under the curves (AUCs) in the range of 0.95–0.99. The highly promising results demonstrate the potential of AI-based computational pathology for aiding pathologists in their diagnostic workflow system. Read full text
Published: 08 Feb 2021 (modified: 21 Apr 2021), Proceedings of Machine Learning Research
Partial transfusion: on the expressive influence of trainable batch norm parameters for transfer learning
Transfer learning from ImageNet is the go-to approach when applying deep learning to medical images. The approach is either to fine-tune a pre-trained model or use it as a feature extractor. Most modern architecture contain batch normalisation layers, and fine-tuning a model with such layers requires taking a few precautions as they consist of trainable and non-trainable weights and have two operating modes: training and inference. Attention is primarily given to the non-trainable weights used during inference, as they are the primary source of unexpected behaviour or degradation in performance during transfer learning. It is typically recommended to fine-tune the model with the batch normalisation layers kept in inference mode during both training and inference. In this paper, we pay closer attention instead to the trainable weights of the batch normalisation layers, and we explore their expressive influence in the context of transfer learning. We find that only fine-tuning the trainable weights (scale and centre) of the batch normalisation layers leads to similar performance as to fine-tuning all of the weights, with the added benefit of faster convergence. We demonstrate this on a variety of seven publicly available medical imaging datasets, using four different model architectures. Read full text
Published: 30 June 2021, Technology in Cancer Research & Treatment
Deep Learning Models for Gastric Signet Ring Cell Carcinoma Classification in Whole Slide Images
Signet ring cell carcinoma (SRCC) of the stomach is a rare type of cancer with a slowly rising incidence. It tends to be more difficult to detect by pathologists, mainly due to its cellular morphology and diffuse invasion manner, and it has poor prognosis when detected at an advanced stage. Computational pathology tools that can assist pathologists in detecting SRCC would be of a massive benefit. In this paper, we trained deep learning models using transfer learning, fully-supervised learning, and weakly-supervised learning to predict SRCC in Whole Slide Images (WSIs) using a training set of 1,765 WSIs. We evaluated the models on two different test sets (n = 999, n = 455). The best model achieved a ROC-AUC of at least 0.99 on all two test sets, setting a top baseline performance for SRCC WSI classification. Read full text
Published: 19 April 2021, Scientific Reports
A deep learning model to detect pancreatic ductal adenocarcinoma on endoscopic ultrasound-guided fine-needle biopsy
Histopathological diagnosis of pancreatic ductal adenocarcinoma (PDAC) on endoscopic ultrasonography-guided fine-needle biopsy (EUS-FNB) specimens has become the mainstay of preoperative pathological diagnosis. However, on EUS-FNB specimens, accurate histopathological evaluation is difficult due to low specimen volume with isolated cancer cells and high contamination of blood, inflammatory and digestive tract cells. In this study, we performed annotations for training sets by expert pancreatic pathologists and trained a deep learning model to assess PDAC on EUS-FNB of the pancreas in histopathological whole-slide images. We obtained a high receiver operator curve area under the curve of 0.984, accuracy of 0.9417, sensitivity of 0.9302 and specificity of 0.9706. Our model was able to accurately detect difficult cases of isolated and low volume cancer cells. If adopted as a supportive system in routine diagnosis of pancreatic EUS-FNB specimens, our model has the potential to aid pathologists diagnose difficult cases. Read full text
Published: 14 April 2021, Scientific Reports
A deep learning model for the classification of indeterminate lung carcinoma in biopsy whole slide images
The differentiation between major histological types of lung cancer, such as adenocarcinoma (ADC), squamous cell carcinoma (SCC), and small-cell lung cancer (SCLC) is of crucial importance for determining optimum cancer treatment. Hematoxylin and Eosin (H&E)-stained slides of small transbronchial lung biopsy (TBLB) are one of the primary sources for making a diagnosis; however, a subset of cases present a challenge for pathologists to diagnose from H&E-stained slides alone, and these either require further immunohistochemistry or are deferred to surgical resection for definitive diagnosis. We trained a deep learning model to classify H&E-stained Whole Slide Images of TBLB specimens into ADC, SCC, SCLC, and non-neoplastic using a training set of 579 WSIs. The trained model was capable of classifying an independent test set of 83 challenging indeterminate cases with a receiver operator curve area under the curve (AUC) of 0.99. We further evaluated the model on four independent test sets—one TBLB and three surgical, with combined total of 2407 WSIs—demonstrating highly promising results with AUCs ranging from 0.94 to 0.99. Read full text
Published: 09 June 2020, Scientific Reports
Weakly-supervised learning for lung carcinoma classification using deep learning
Lung cancer is one of the major causes of cancer-related deaths in many countries around the world, and its histopathological diagnosis is crucial for deciding on optimum treatment strategies. Recently, Artificial Intelligence (AI) deep learning models have been widely shown to be useful in various medical fields, particularly image and pathological diagnoses; however, AI models for the pathological diagnosis of pulmonary lesions that have been validated on large-scale test sets are yet to be seen. We trained a Convolution Neural Network (CNN) based on the EfficientNet-B3 architecture, using transfer learning and weakly-supervised learning, to predict carcinoma in Whole Slide Images (WSIs) using a training dataset of 3,554 WSIs. We obtained highly promising results for differentiating between lung carcinoma and non-neoplastic with high Receiver Operator Curve (ROC) area under the curves (AUCs) on four independent test sets (ROC AUCs of 0.975, 0.974, 0.988, and 0.981, respectively). Development and validation of algorithms such as ours are important initial steps in the development of software suites that could be adopted in routine pathological practices and potentially help reduce the burden on pathologists. Read full text
Published: 30 January 2020, Scientific Reports
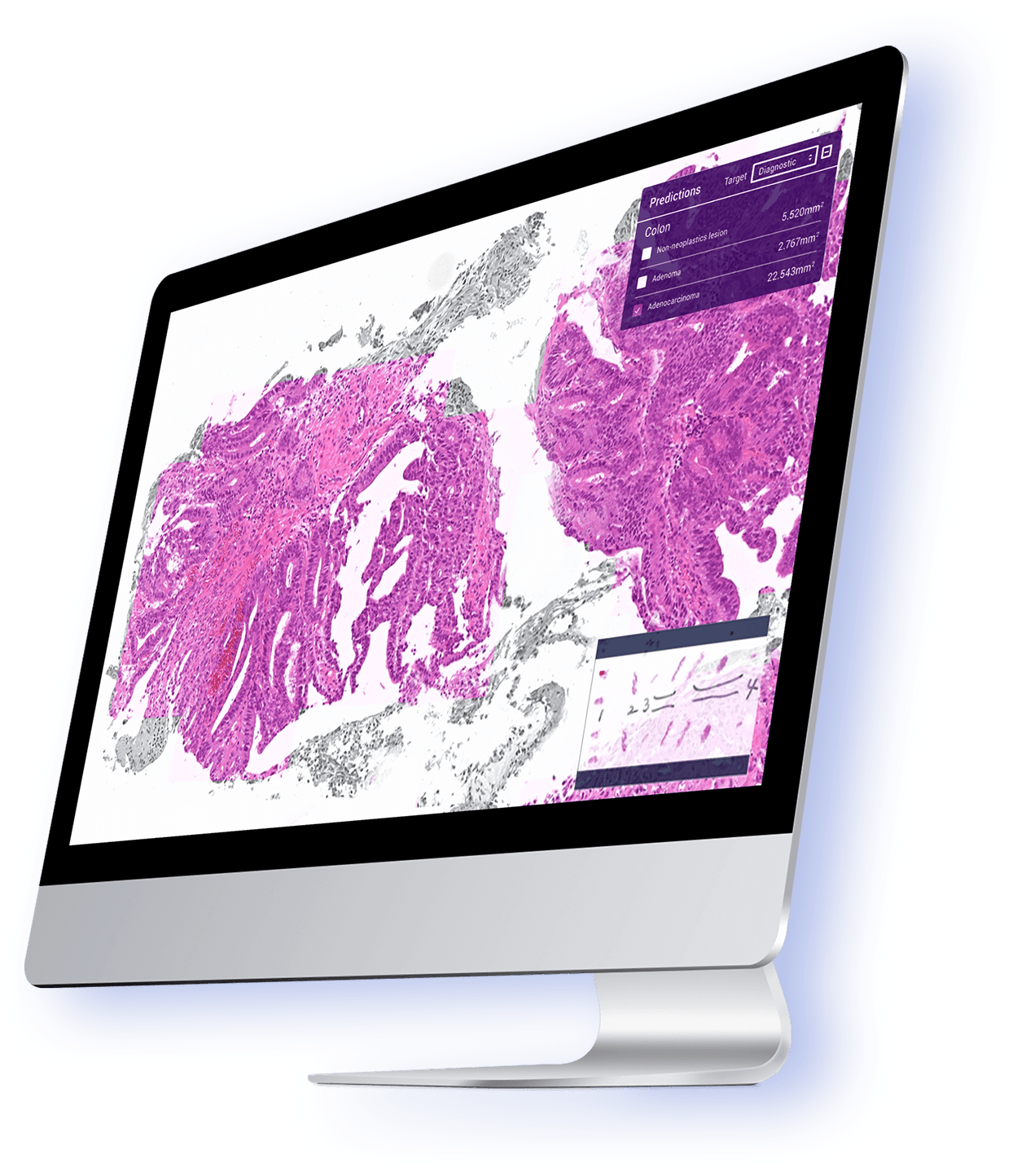
Deep Learning Models for Histopathological Classification of Gastric and Colonic Epithelial Tumours
Histopathological classification of gastric and colonic epithelial tumours is one of the routine pathological diagnosis tasks for pathologists. Computational pathology techniques based on Artificial intelligence (AI) would be of high benefit in easing the ever increasing workloads on pathologists, especially in regions that have shortages in access to pathological diagnosis services. In this study, we trained convolutional neural networks (CNNs) and recurrent neural networks (RNNs) on biopsy histopathology whole-slide images (WSIs) of stomach and colon. The models were trained to classify WSI into adenocarcinoma, adenoma, and non-neoplastic. We evaluated our models on three independent test sets each, achieving area under the curves (AUCs) up to 0.97 and 0.99 for gastric adenocarcinoma and adenoma, respectively, and 0.96 and 0.99 for colonic adenocarcinoma and adenoma respectively. The results demonstrate the generalisation ability of our models and the high promising potential of deployment in a practical histopathological diagnostic workflow system. Read full text